-Search query
-Search result
Showing 1 - 50 of 65 items for (author: zhou & jm)

EMDB-41374: 
Antibody N3-1 bound to RBDs in the up and down conformations
Method: single particle / : Hsieh CL, McLellan JS
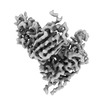
EMDB-41382: 
Antibody N3-1 bound to RBD in the up conformation
Method: single particle / : Hsieh CL, McLellan JS
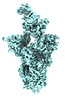
EMDB-41399: 
Antibody N3-1 bound to SARS-CoV-2 spike
Method: single particle / : Hsieh CL, McLellan JS

EMDB-41370: 
Structure of a class A GPCR/Fab complex
Method: single particle / : Sun D, Johnson M, Masureel M

EMDB-41827: 
Structure of a class A GPCR/agonist complex (focused map2)
Method: single particle / : Sun D, Johnson M, Masureel M

EMDB-41828: 
Structure of a class A GPCR/agonist complex (focused map1)
Method: single particle / : Sun D, Johnson M, Masureel M

EMDB-41829: 
Structure of a class A GPCR/agonist complex
Method: single particle / : Sun D, Johnson M, Masureel M

EMDB-41850: 
Structure of a class A GPCR/agonist complex (Consensus map)
Method: single particle / : Sun D, Johnson M, Masureel M

EMDB-41144: 
Cryo-EM Structure of GPR61-G protein complex stabilized by scFv16
Method: single particle / : Lees JA, Dias JM, Han S

EMDB-41145: 
Cryo-EM Structure of GPR61-
Method: single particle / : Lees JA, Dias JM, Han S

EMDB-34710: 
Cryo-EM structure of WeiTsing
Method: single particle / : Qin L, Tang LH, Chen YH

EMDB-40094: 
17b10 fab in complex with full-length SARS-CoV-2 Spike G614 trimer
Method: single particle / : Kwon HJ, Zhang J, Kosikova M, Tang WC, Rodriguez UO, Peng HQ, Meseda CA, Pedro CL, Schmeisser F, Lu JM, Zhou B, Davis CT, Wentworth DE, Chen WH, Shriver MC, Pasetti MF, Weir JP, Chen B, Xie H

EMDB-40095: 
17B10 fab in complex with up-RBD of SARS-CoV-2 Spike G614 trimer
Method: single particle / : Kwon HJ, Zhang J, Kosikova M, Tang WC, Rodriguez UO, Peng HQ, Meseda CA, Pedro CL, Schmeisser F, Lu JM, Zhou B, Davis CT, Wentworth DE, Chen WH, Shriver MC, Pasetti MF, Weir JP, Chen B, Xie H

EMDB-13895: 
CryoEM structure of the Smc5/6-holocomplex (composite structure)
Method: single particle / : Hallett ST, Oliver AW

EMDB-13893: 
Cryo-EM structure of the Smc5/6 holo-complex; map for head-end of complex.
Method: single particle / : OLIVER AW, Hallett ST

EMDB-13894: 
CryoEM structure of the Smc5/6 holocomplex; map for hinge and arm region.
Method: single particle / : OLIVER AW, Hallett ST

EMDB-26522: 
SARS-CoV-2 6P Mut7 in complex with K398.25 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26523: 
SARS-CoV-1 in complex with K398.25 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26524: 
SARS-CoV-2 6P Mut7 in complex with K398.16 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB
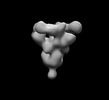
EMDB-26525: 
SARS-CoV-2 6P Mut7 in complex with K398.16 Fab (3 bound)
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26526: 
SARS-CoV-1 in complex with K398.16 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26527: 
SARS-CoV-2 6P Mut7 in complex with K288.2 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26528: 
SARS-CoV-2 6P Mut7 in complex with K398.8 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26529: 
SARS-CoV-2 6P Mut7 in complex with K398.8 Fab (2 bound)
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26530: 
SARS-CoV-2 6P Mut7 in complex with K398.18 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26531: 
SARS-CoV-2 6P Mut7 in complex with K398.18 Fabs (2 bound)
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26532: 
SARS-CoV-2 6P Mut7 in complex with K398.22 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB
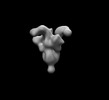
EMDB-26533: 
SARS-CoV-2 6P Mut7 in complex with K398.22 Fab (2 bound)
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26534: 
SARS-CoV-1 in complex with K398.8 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26535: 
SARS-CoV-1 in complex with K398.8 Fab (2 bound)
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26536: 
SARS-CoV-1 in complex with K398.18 Fab (2 bound)
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26537: 
SARS-CoV-1 in complex with K398.18 Fab (3 bound)
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26538: 
SARS-CoV-1 in complex with K288.2 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-26539: 
SARS-CoV-1 in complex with K398.22 Fab
Method: single particle / : Lee WH, Torres JL, Ward AB

EMDB-12957: 
Cryo-EM structure of the human R2TP-TTT complex
Method: single particle / : Pal M, Llorca O, Pearl L

EMDB-12979: 
Cryo-EM structure of the TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex
Method: single particle / : Pal M, Llorca O, Pearl L

EMDB-22943: 
Cryo-EM structure of single ACE2-bound SARS-CoV-2 trimer spike at pH 5.5
Method: single particle / : Gorman J, Rapp M, Kwong PD, Shapiro L

EMDB-22949: 
Cryo-EM Structure of Double ACE2-Bound SARS-CoV-2 Trimer Spike at pH 5.5
Method: single particle / : Gorman J, Rapp M, Kwong PD, Shapiro L

EMDB-22950: 
Cryo-EM structure of Triple ACE2-bound SARS-CoV-2 Trimer Spike at pH 5.5
Method: single particle / : Gorman J, Rapp M, Kwong PD, Shapiro L

EMDB-22922: 
ACE2-RBD Focused Refinement Using Symmetry Expansion of Applied C3 for Triple ACE2-bound SARS-CoV-2 Trimer Spike at pH 7.4
Method: single particle / : Gorman J, Kwong PD, Shapiro L

EMDB-22927: 
Cryo-EM structure of triple ACE2-bound SARS-CoV-2 trimer spike at pH 7.4
Method: single particle / : Gorman J, Kwong PD, Shapiro L

EMDB-22932: 
Cryo-EM structure of double ACE2-bound SARS-CoV-2 trimer Spike at pH 7.4
Method: single particle / : Gorman J, Kwong PD, Shapiro L

EMDB-22941: 
Cryo-EM structure of single ACE2-bound SARS-CoV-2 trimer spike at pH 7.4
Method: single particle / : Gorman J, Kwong PD, Shapiro L

EMDB-22515: 
Structure of SARS-CoV-2 spike at pH 4.5
Method: single particle / : Tsybovsky Y, Zhou T, Kwong PD

EMDB-30476: 
Cryo-EM structure of Chikungunya virus in complex with Fab fragments of mAb CHK-124
Method: single particle / : Zhou QF, Fox JM, Earnest JT, Ng TS, Kim AS, Fibriansah G, Kostyuchenko VA, Shu B, Diamond MS, Lok SM

EMDB-30477: 
Cryo-EM structure of Chikungunya virus in complex with Fab fragments of mAb CHK-263
Method: single particle / : Zhou QF, Fox JM, Earnest JT, Ng TS, Kim AS, Fibriansah G, Kostyuchenko VA, Shu B, Diamond MS, Lok SM

EMDB-30478: 
Cryo-EM structure of Chikungunya virus in complex with mAb CHK-263 IgG
Method: single particle / : Zhou QF, Fox JM, Earnest JT, Ng TS, Kim AS, Fibriansah G, Kostyuchenko VA, Shu B, Diamond MS, Lok SM

EMDB-30479: 
Cryo-EM structure of Chikungunya virus in complex with Fab fragments of mAb CHK-263 (subregion around icosahedral 5-fold vertex)
Method: single particle / : Zhou QF, Fox JM, Earnest JT, Ng TS, Kim AS, Fibriansah G, Kostyuchenko VA, Shu B, Diamond MS, Lok SM

EMDB-30480: 
Cryo-EM structure of Chikungunya virus in complex with mAb CHK-263 IgG (subregion around icosahedral 2-fold vertex)
Method: single particle / : Zhou QF, Fox JM, Earnest JT, Ng TS, Kim AS, Fibriansah G, Kostyuchenko VA, Shu B, Diamond MS, Lok SM

EMDB-22829: 
Human Tom70 in complex with SARS CoV2 Orf9b
Method: single particle / : QCRG Structural Biology Consortium
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
